First SARS-CoV-2 standard designed specifically for NGS applications
The problem:
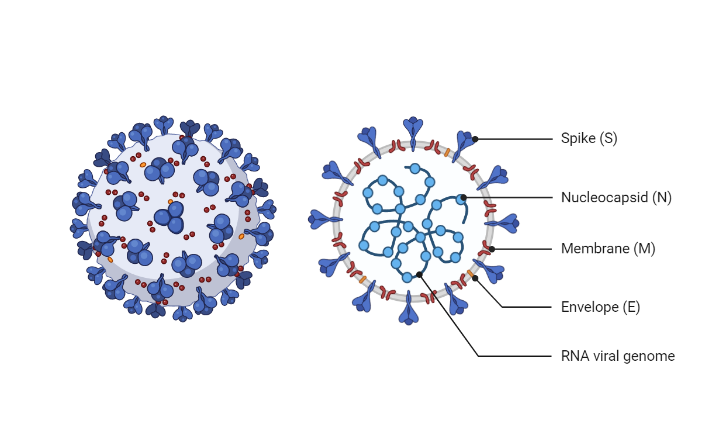
While there are several synthetic reference materials for use with analytical validation for multiple molecular methods to detect SARS-CoV-2, there are no NGS-specific run standards. SARS-CoV-2 can be challenging to sequence and ensure high accuracy of sequence assembly and correct identification of variants. Moreover, there are no viral specific internal controls that help to standardize the workflow and measure the performance of the reagents.
The product:
The Accukit SARS-CoV-2 RNA standard is a mixture of 17 synthetic RNA overlapping constructs spanning the entire SARS-CoV-2 genome. When added to the RNA from the extracted sample prior to the reverse transcription (RT) step, will control for systemic and technical errors through the entire workflow. It also contains a complexity control (65k unique molecules) to estimate library preparation and bioinformatic impact on complexity capture for each sample.
Why does it help?
SEQUENCE ACCURACY (LIMIT OF BLANK)
Accukit SARS-CoV-2 RNA standards co-located with the target sample measure background error due to library prep, sequencing, or other factors. The input sequence of the SARS-CoV-2 standards are known so any output variation would have arisen from a process error. Thus, the standards provide a limit of blank (LOB) for every position measured and help to eliminate variant caller error, particularly with low-frequency alleles. The complexity control provides an independent quantitative measure of each sample’s molecular diversity yield in support of assessing reagent quality and performance drift.
LIMIT OF DETECTION (LOD)
When added in a limiting amount to every sample, Accukit SARS-CoV-2 provides LOD control for qualitative detection assays. By virtue of being in each sample, it assures the assay met its sensitivity performance threshold, even in the absence of a positive target. A positive control for a negative result.
RUN QC
Standardize intra- and inter-lab testing, across platforms, operators, assays, and geographies to improve the quality of the result and ensure optimal reagent performance with every run.
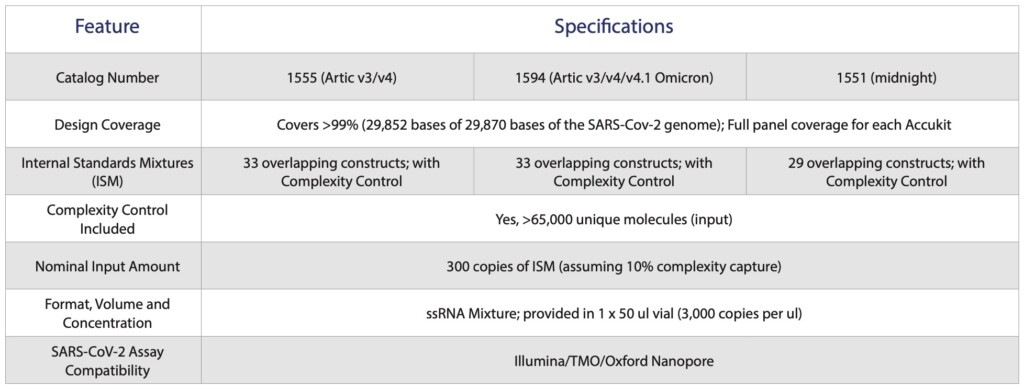
Download the product sheet below.
